Human studies
All subjects gave their written consent to take part in the experiment, executed at the Copenhagen University Department for Nutrition, Exercise and Sports, approved by the Copenhagen Ethics Committee (H-18005023) and registered in a public database, Clinical Trials.gov (NCT06267235).
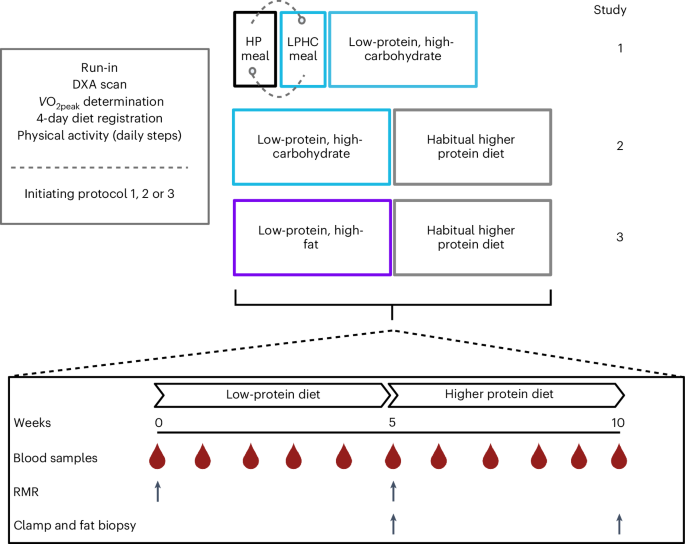
Three studies were performed in three groups of individuals (Fig. 1).
Study 1, acute and prolonged reduced protein intake replaced by carbohydrates
To evaluate the acute postprandial metabolic effects, ingestion of a LPHC meal was compared with a meal with a HP content (Fig. 1), which was equivalent to the participants’ habitual protein intake. Eight healthy, young (age 27 ± 2 years), lean (body mass index (BMI), 25 ± 2 kg m−2), moderately physically active (peak oxygen uptake (VO2peak), 44 ml min−1 kg−1 body mass) men (Table 1) with a daily protein intake of ≥1.5 g kg−1 body weight were recruited. The LPHC and HP meal tests were performed in a randomized order, separated by 72 h. The LPHC meal consisted of 8 E% protein, 71 E% carbohydrate and 21 E% fat; the HP meal consisted of 16 E% protein, 57 E% carbohydrate and 27 E% fat (Extended Data Table 1) and reflected the habitual dietary macronutrient composition of the participants. After arrival at the institute in the morning in an overnight-fasted state (10 h), a catheter was inserted into an antecubital vein for blood sampling. Then the participants ingested the meal within 10 min, together with 200 ml of water. The energy intake amounted to 55 KJ kg−1 body weight. Blood samples were drawn before (0 min) the meal and at 15, 30, 45, 60, 90, 120, 180 and 240 min after ingestion. Indirect calorimetry was applied at 0, 30, 60, 90, 120, 180 and 240 min. After the second meal test, the participants continued a eucaloric diet reduced in protein (0.8 g kg−1 body weight per day) replaced by carbohydrates (LPHC) for 5 weeks (9 E% protein, 70 E% carbohydrate and 21 E% fat) (Fig. 1 and Extended Data Table 1).
Study 2, protein-reduced diet substituted with carbohydrates for 5 weeks followed by 5 weeks on habitual HPD
Eight healthy, young (age, 26 ± 2 years), lean (BMI, 25 ± 3 kg m–2), moderately physically active (VO2peak, 44 ± 5 ml min−1 kg−1) men (Table 1), with a daily protein intake of ≥1.5 g kg−1 body weight were recruited to this study. The participants consumed the eucaloric LPHC as described in study 1 (Extended Data Table 2) for 5 weeks, followed by another 5 weeks on a eucaloric diet consisting of the participants’ habitual HPD (18 E% protein, 49 E% carbohydrate, 33 E% fat) (Fig. 1 and Table 2).
Study 3, protein-reduced diet substituted with fat for 5 weeks followed by 5 weeks on the habitual HPD
Seven healthy, young (age, 25 ± 1 years), lean (BMI, 23 ± 2 kg m−2), moderately physically active (VO2peak, 41 ± 8.0 ml min−1 kg−1) men (Table 1), with a daily protein intake of ≥1.5 g kg−1 body weight matching the participants in study 1 and 2 were enrolled. One participant dropped out because of illness. The participants consumed a eucaloric LPHF diet, consisting of 9 E% protein, 41 E% carbohydrate and 50 E% fat for 5 weeks, followed by another 5 weeks on their habitual HPD (18 E% protein, 49 E% carbohydrate, 33 E% fat) (Fig. 1 and Table 3).
Experimental diets and eucaloric weight maintenance
In a run-in period of at least 1 week before each of the dietary interventions (Fig. 1), energy content and macronutrient composition of the participants’ habitual diet were evaluated from a weighed dietary registration for four non-consecutive days (Extended Data Table 3). All food and fluids were weighed to an accuracy of 1 g and analysed with computer software (Vitakost, Denmark). The daily energy provision on the experimental diets was based on the individual diet registrations adjusted based on previously published basal metabolic rate equations47 and multiplied by the physical activity level value, estimated from fitness level (VO2peak), measured on a bicycle ergometer and pediometrics and accelerometrics. To maintain the baseline body weight of the participants throughout the interventions, participants registered their morning weight daily. Energy provision was adjusted during the diet intervention if body weight, based on a 3-day average, changed by ±0.5 kg. All foods in the experimental diets were weighted to 1 g of accuracy and pre-packed in meal portions in a 7-day menu rotation, and food was picked up by participants every third day at the institute. The LPHC diet contained a high proportion of carbohydrates derived from white bread, rice, pasta and fruits. The LPHF diet contained a high proportion of fat derived from vegetable oil, nuts, butter, cheese, milk and yoghurt with high fat content. The HPD reflected the habitual mixed diet of the participants.
Experimental study
Before and after the LPHC diet (study 1) and before and weekly during the LPHC and LPHF diets as well as the HPD (studies 2 and 3, Fig. 1), participants arrived in the morning at the institute by passive transport in an overnight-fasted state (10 h). After resting in the supine position for at least 30 min under thermoneutral conditions, RMR was measured by indirect calorimetry (Masterscreen CPX SBX, CareFusion) at week 0 and week 5 (studies 2 and 3). Then a catheter was inserted into the antecubital vein from which blood samples were obtained. In study 1, body composition was assessed using DXA-scanning (Lunar Corporation) before and at week 5. In studies 2 and 3, body composition was evaluated before, at week 5 and at week 10. Additionally, at weeks 5 and 10, body water content was measured by bio-impedance (InBody270).
Whole-body insulin sensitivity
Participants in studies 2 and 3 underwent a hyperinsulinemic-euglycemic clamp at the end of week 5 of the LPHC, LPHF and HPD diet (Fig. 1). Participants arrived at the institute in the morning after an overnight fast (10 h) by passive transport. A catheter was inserted in the antecubital vein and fasting blood samples were drawn for basal measurements and 2D glucose background enrichment. Then, a bolus injection of [6,6-D2] glucose tracer (2.6 mg kg−1) was given, followed by constant infusion (0.044 mg kg−1 min−1) to determine basal hepatic glucose production. Another catheter was inserted into a dorsal hand vein in the other arm, and a heating pad was wrapped around the hand to arterialize the venous blood from the hand. After 120 min tracer infusion, a 120 min hyperinsulinemic-euglycemic clamp was initiated by a bolus of insulin (9.0 mU kg−1) (Actrapid, Novo Nordisk) followed by a constant infusion rate (1.0 mU insulin per kg per min). During the clamp, 20% glucose solution enriched with 1.9% [6.6-D2]-glucose tracer was infused at a rate ensuring euglycemia, matching the fasting arterialized blood glucose level determined from three blood samples obtained before initiating the insulin infusion. The same glucose target was used after the HPD. Blood samples were drawn before (0 min) and at 30, 90 and 120 min during the clamp, and indirect calorimetry was applied at 0, 45, 60 and 120 min. Biopsies were obtained from the periumbilical subcutaneous adipose tissue by a modified Bergström needle with suction under local anaesthesia (∼2 ml of xylocaine 1%; AstraZeneca) before the clamp. Biopsies were rinsed in ice-cold saline, snap-frozen in nitrogen and stored at −80 °C until analysis. Clamp data from two subjects (LPHC intervention, study 2) were excluded: one subject fainted after the basal biopsy and one was excluded because of technical problems during the clamp. Data on hepatic glucose production are missing from one subject (LPHF, study 3) because of a lack of glucose tracer.
Physical activity
Participants were instructed to maintain their physical activity level during each intervention. Before and after the LPHC, LPHF and HPD interventions, maximal oxygen uptake (VO2peak) was measured (Masterscreen CPX SBX, CareFusion) using an incremental test to exhaustion on a bicycle ergometer. To monitor daily physical activity levels during the interventions, participants wore either a pedometric watch (Polar Loop-watch or a Garmin Vivofit3-watch; study 1) or a triaxial accelerometer (SENS motion system, Denmark; study 2 and 3) attached to the skin on the lateral thigh approximately 10 cm from the knee to measure daily number of steps and activities.
Plasma analysis
See resource table (Extended Data Table 5).
Calculations
The HOMA-IR index was calculated as (insulinfasting) × (glucosefasting) / 22.5.
Hepatic glucose production in the fasting state and during insulin stimulation was calculated from three blood samples obtained during the last 20 min of the basal period and again during the last 20 min of the clamp period using Steele’s equation, taking into account both the ‘cold’ glucose concentration and the enrichment of glucose tracer in the blood during these periods as previously described48. RMR was calculated based on a previous publication49 as EE (kJ min−1) = VO2 × (4.686 + (RQ − 0.707) × 0.361 / 0.293) × 4.14 kJ kcal−1, where EE is energy expenditure and RQ is the respiratory quotient.
Animal study
Animal handling and experimentation was done at the German Cancer Research Centre (Heidelberg) in accordance with European Union directives and the German Animal Welfare Act and approved by local authorities (Regierungspräsdidium Karlsruhe) and conformed to ARRIVE guidelines. WT C57BL/6J male mice were obtained from Charles River Laboratories. All mice were maintained on a 12 h light–dark cycle (06:00–18:00 h) at 22 °C with unrestricted access to food and water. Germline FGF21-KO male mice were generated on a C57BL/6 background as previously described14. FGF21+/− mice were crossed by het × het littermate pairing to generate FGF21-KO (−/−) and WT (+/+) littermates, bred at the German Cancer Research Center. All experimental procedures have been described in detail elsewhere14. In short, 8-week-old mice were housed with two to three mice per cage and fed ad libitum with either a LPHC diet (5 E% protein and 85 E% carbohydrate; Research Diets, D10062201) or a LPHF diet (5 E% protein and 60 E% fat; Research Diets, D12020703), with representative mice fed on a standard HPD (protein 20 E%; Research Diets, D12450B and D12492). Mice were weighed before allocation to diet groups by counterbalancing. Food intake and body weight were measured before and after the diet interventions. After 10 weeks, mice were killed and iWAT was dissected and snap-frozen in nitrogen. No animals were excluded from the analysis.
Adipose tissue proteomics and bioinformatics
A total of 50 mg adipose tissue in 5× volume lysis buffer (2% sodium deoxycholate, 100 mM Tris pH 8.5) was boiled for 10 min at 99 °C. After homogenization with pestle, samples were sonicated for 15 min (Branson probe sonifier, output 3–4, 50% duty cycle, 15 min with 30 s × 30 s cycles), and protein content was determined with the bicinchoninic acid method (BCA no. 23225, Pierce) and adjusted to a protein level at 0.25 µg µl−1. Then, proteins were alkylated (40 mM 2-chloroacetamide and 10 mM trifluoroacetic acid (TFA)) in the dark for 10 min at 45 °C at 1,000 rpm in a thermoshaker and digested with LysC and trypsin (1:50 protein to enzyme) overnight at 37 °C, 1,000 r.p.m. in a thermoshaker. Digested peptides were acidified by adding 1:1 isopropanol with 1% TFA and loaded in triple layer Styrene Divinylbenzene–Reversed Phase Sulfonate STAGE tips (SDB–RPS; 3M Empore). The STAGE tips were first cleaned (100 µl 80% acetonitrile (ACN)) then activated (100 µl 30% methanol + 1% TFA) and acidified (150 µl 0.2% TFA). The peptides were eluted with 60 µl SDB–RPS elution buffer (80% ACN, 5% NH4OH) on a SpeedVac for 45 min at 45 °C and dissolved in 6 µl MS loading buffer (2% ACN, 0.01% TFA).
Proteomic samples were measured at a Thermo Exploris 480 combined with a Thermo EasyNLC 1200 and Thermo FAIMS Pro using a 60 min method with single collision voltage of −50 V in data-independent acquisition (DIA) mode. The input material for each sample was 500 µg of peptides, based on Nanodrop measurement.
Liquid chromatography gradients were provided by incrementally mixing buffer A (0.1% formic acid) and buffer B (80% acetonitrile, 0.1% formic acid). After starting with 5% buffer B, the amount was linearly increased to 20% after 30 min, 29% after 39 min, 45% after 45 min and 95% after 50 min, followed by holding 95% for another 5 min (until min 55) and finally reduced to 5% after 60 min at a flow of 300 nl min−1.
MS1 scans were acquired with an orbitrap resolution of 120,000 in positive ion mode. HCD collision energy was 30%. For MS2 scans, an orbitrap resolution of 15,000 was used with 66 DIA windows of variable size.
Statistical analyses
Data collection and analysis were not performed blind to the conditions of the experiments. Statistical analyses were performed in GraphPad Prism (v.8 and v.9). Data distribution was assumed to follow the Gaussian normal distribution as well as variance homogeneity, but this was not formally tested. To assess differences between interventions and/or the effect of time, repeated measures one-way or two-way ANOVA was performed, or a mixed-effects model in the case of missing values. When ANOVA revealed significant interactions, Bonferroni post hoc testing with correction for multiple testing was applied. When comparing pre-intervention versus post-intervention or comparing between groups, a two-tailed paired t-test or unpaired t-test was used. To assess the relationship between changes in energy intake and circulating FGF21 levels, a repeated measures correlation was conducted using percentage delta values from baseline to week 5 and from week 5 to week 10 among participants enrolled in studies 2 and 3. This method addresses the issue of non-independence among observations, providing a common within-individual association. The analysis was conducted using the rmcorr package in R (v.4.3.3) with RStudio. For proteome analyses, the raw data were analysed using Spectronaut (Biognosis) in directDIA mode, using a library-free approach. All searches were performed against the human UniProt FASTA database, with MaxLFQ settings enabled for protein-level label-free quantification. The MaxQuant contaminant FASTA file was included in the search, and default parameters were retained unless explicitly stated otherwise. Bioinformatics analysis was conducted using Perseus, applying default settings unless specified. Before analysis, common contaminants were filtered out. For total proteome analysis, LFQ values were transformed to a logarithmic scale (log2), and proteins were selected if they had at least two valid measurements across biological replicates for any given condition. Missing values were imputed based on a Gaussian normal distribution model with a width of 0.3 and a downshift of 1.8. Differentially regulated proteins between two conditions were identified using an unadjusted two-sided Student’s t-test. No statistical methods were used to pre-determine sample sizes but our sample sizes are similar to those reported in previous publications14. All data are presented as mean ± s.e.m. except for subject characteristics in Table 1 (mean ± s.d.). For all statistical tests, P < 0.05 was considered statistically significant.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
link



:max_bytes(150000):strip_icc()/GettyImages-1070405846-1f2cbced7968419a8bcc9c7115c1e09c.jpg)



