Data description
For the sake of robustness of analyses, we modelled the performance of world-class short-track speed skaters over two distinct datasets, collected in 2019 and 2016 respectively. For both datasets, participants were fully informed about data collection and written consent was obtained from them. Also, the study was performed in agreement with the standards set by the declaration of Helsinki (2013) involving human subjects. The protocol was reviewed and approved by the local research Ethics Committee (EuroMov, University of Montpellier, France). The present retrospective study relied on the collected data without causing any changes in the training programming of athletes.
The 2019 dataset
Nine (9) national elite short-track speed skaters were recruited in a previous study12. Two (2) athletes were injured during the training range and were discarded from the study. Details about the population of interest (i.e. 7 athletes) are provided by Imbach, Perrey, et al.12.
The data collection spread over 10 weeks, from June to September. The period of study was interrupted by a two-week summer break, during which athletes fully rested. Skaters sustained a mean weekly training volume of 16.6 ± 2.5 h during this period. A total of 375 training and 248 performance observations were collected, distributed as 53.6 ± 1.6 training observations and 35.4 ± 2.2 performances per athlete.
The performance data were collected each week and consisted in the time to perform a 166.68 m race after a standardised warm-up; hence a lower performance value was sought. Timing gate systems (Brower timing system, USA) were used to record valid individual time trial performance17. On-ice and off-ice training loads were used as the explanatory variable as described in the Appendix 1 from Imbach, Perrey, et al.12.
The 2016 dataset
Fifteen (15) national elite Short-track speed skaters voluntary participated to a previous study18. One of the skaters was injured and removed from the study, leaving 14 athletes in. Athletes had the same coach for both datasets. Out of these 14 athletes, 2 were also represented in the 2019 data set.
A total of 948 training and 755 performance observations were collected, distributed as 67.7 ± 5.6 training observations and 53.9 ± 5.4 performances per athlete. The period of study included 13 weeks from July to October. During this period, the skaters sustained a mean weekly training volume of 25 ± 5 h.
Following a 15-minutes standardised warm-up, participants performed weekly flying starts for a one-lap maximal run. Four photocells were settled for recording skating times. The dependent variable further modelled was the individual maximal acceleration (from speed derivation); hence a higher performance value was sought. On-ice and off-ice training sessions were used as the explanatory variables, as described by Méline et al.18 and consistently with the 2019 dataset.
Models
We considered the Banister’s fitness-fatigue model (FFM) from a stochastic perspective, in which the predicted performance of athlete s at time t: \(\:{y}_{s}\left(t\right)\), was considered as the expectation from a gaussian distribution:
$$\:{y}_{s}\left(t\right)\sim\:N({\widehat{y}}_{s}\left(t\right),\:{\sigma\:}_{s}^{2})$$
(1)
of variance \(\:{\sigma\:}_{s}^{2}\) and expectation \(\:{\widehat{y}}_{s}\left(t\right)\) a function of the daily training doses or workloads \(\:{w}_{s}\left(i\right)\)1:
$$\:{\widehat{y}}_{FFM,\:\:s}\left(t\right)=\:{a}_{0,s}+{k}_{G,s}\sum\:_{i=1}^{t-1}{w}_{s}\left(i\right){e}^{\frac{-(t-i)}{{\tau\:}_{G,s}}}\:-{k}_{H,s}\sum\:_{i=1}^{t-1}{w}_{s}\left(i\right){e}^{\frac{-(t-i)}{{\tau\:}_{H,s}}}$$
In the above equation, a0 denotes the intercept, kG and kH represent the linear coefficients for fitness and fatigue, respectively, and τG and τH the time exponents for fitness and fatigue, respectively. All these parameters are written with an s subscript that denotes the athlete identity, as the parameters are estimated at the individual level.
The FFM being relatively complex, we decomposed it into a simpler, potentially more parsimonious version. This simpler version only encompassed a fitness component:
$$\:{\widehat{y}}_{FOM,s}\left(t\right)=\:{a}_{0,s}+{k}_{G,s}\sum\:_{i=1}^{t-1}{w}_{s}\left(i\right){e}^{\frac{-(t-i)}{{\tau\:}_{G,s}}}$$
This is referred to as the fitness-only model (FOM) in this study.
Model fitting
Prior knowledge on the time exponents
One of the main interests of the FFM is its biological interpretability. Exponential decays can be interpreted from their half-life (i.e. the amount of time needed for half the effect of an input to vanish). The time exponent value τ leading to a half-life h is:
$$\:{e}^{\frac{-x}{\tau\:}}=0.5\iff\:\:\:\tau\:=\frac{h}{\text{l}\text{n}\left(2\right)}$$
In this study, we investigated the option to provide information to τG and τH either by fixing them in a frequentist framework or using bayesian priors. Priors were based on background, prior knowledge that an athlete may recover half the fatigue of a training session in about the length of a weekend (i.e. 2 days) and may lose half of the fitness from a training session in about a month (i.e. 30 days). Therefore, when informing the time exponents, we provided the target value of τG = 43.28 for fitness and τH = 2.89 for fatigue.
Frequentist model fitting
The frequentist model fitting refers to the estimation of θ by minimizing the standard deviation σ, which is equivalent to maximizing the likelihood under Gaussian dispersion assumption (Eq. 1). The minimization was performed using the Powell algorithm from the scipy library19.
The frequentist fitting process consisted in 10 independent runs from which we retained runs that converged and minimized the residual standard deviation. Since the model was ill-conditioned, as we explored in depth throughout this study, the parameters estimation became unstable even under constraints optimization (i.e. boundaries) for convergence (Table 1).
Alternatively, we fixed τG and τH (see Sect. 2.3.1) before estimating the other parameters, hence reducing the system to a linear model.
Bayesian model fitting
Models were also fitted in a bayesian framework. We used proper, mildly informative priors for \(\:{a}_{0}\sim\:N(0,{5}^{2})\) ; \(\:{k}_{G}=-\left|X\right|\) (for the 2019 dataset) (or \(\:{k}_{G}=+\left|X\right|\) for the 2016 dataset), with \(\:X\sim\:N(0,{5}^{2})\); idem for \(\:{k}_{H}\) ; and a flatter prior for \(\:{\upsigma\:}\:=\:\left|X\right|\) with \(\:X\sim\:N(0,{25}^{2})\). Half-normal distribution is part of the half-Student distribution family, which are generally advised for variance parameters priors20.
We compared two strategies for the time exponent estimations. On the one hand, we used non-informative priors \(\:{\tau\:}_{G}\sim\varGamma\:(a=1,\:b=0.5)\) which is equivalent to a \(\:{{\upchi\:}}^2\) distribution with 2 degrees of freedom; same for \(\:{\tau\:}_{H}\). On the other hand, we fitted the models with informative priors for the time exponents: \(\:{\tau\:}_{G}\sim\varGamma\:(a=2*43.28,\:b=2)\) and \(\:{\tau\:}_{H}\sim\varGamma\:(a=2*2.89,\:b=2)\). The average of a gamma distribution defined by its shape (a) and rate (b) is a/b, hence these priors are consistent with the background knowledge target as developed in 2.3.1. We emphasize that these targets have been designed without information from the data. The rate parameter b = 2 was set high but not extreme.
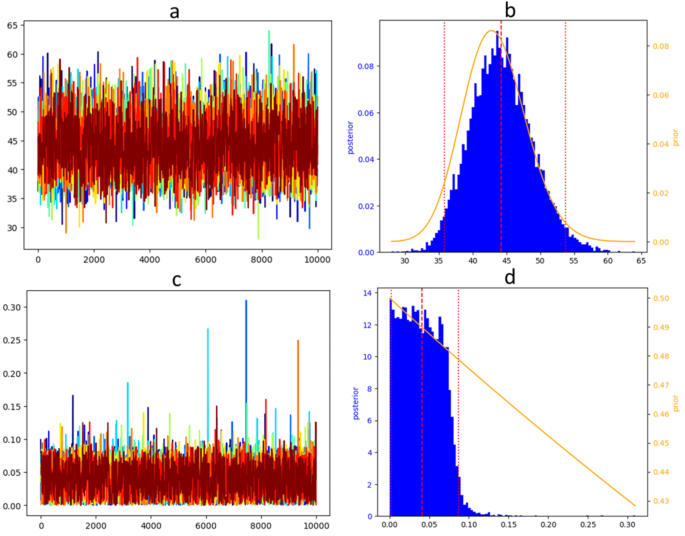
Bayesian fitting was performed using Markov Chain Monte Carlo (MCMC), with 10 independent Markov chains for each model fitting. Independent Markov chains were initiated from random initial values uniformly sampled within the intervals described in Sect. 2.3.2. Each chain had 4 × 104 iterations warm-up, 1 × 104 iterations sampling, and a 10-iteration thinning so they ended up 103 iterations long. Computation were performed using the Python implementation of Stan21.
Cross-validation
The predictive ability of the models was assessed by the mean of cross-validation. Since we deal with time series, the data were time-ordered and split accordingly in subsets in respect of hidden-value block cross-validation with an incremental window12,22. We used minimal values of 35 days and 15 days for training and test sets, respectively. Due to potential dependencies between training and test data, a 5-day gap between each training and test subsets was considered to ensure unbiased model predictions. This parameterization yielded 2 folds for the 2019 dataset and 3 folds for the 2016 dataset as described in Table 2.
We used root mean square error (RMSE) between predicted and observed performance to quantify the predictive ability of the models. RMSE of the training data, i.e. the residual deviation of the model σ, was also employed to measure the goodness of fit of a model to its training data.
link





